-Search query
-Search result
Showing 1 - 50 of 3,688 items for (author: xiao & p)

EMDB-44551: 
Map of eastern equine encephalitis virus q3 spike protein in complex with VLDLR without masked refinement
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-36577: 
Structure of human TRPV1 in complex with antagonist
Method: single particle / : Fan J, Lei X

EMDB-38161: 
Structure of human TRPV1 in complex with antagonist --protein purified without CHS
Method: single particle / : Fan J, Lei X

EMDB-42050: 
Structure of eastern equine encephalitis virus VLP in complex with VLDLR LA1
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-42055: 
Structure of eastern equine encephalitis virus VLP unliganded quasi-threefold spike protein
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.
Method: single particle / : Acharya P, Parsons R, Janowska K, Williams WB, Alam M, Haynes BF
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
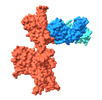
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
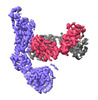
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
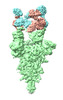
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
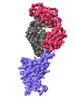
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-45078: 
E.coli GroEL + PBZ1587 inhibitor
Method: single particle / : Watson ER, Lander GC
EMDB-45079: 
E.coli GroEL apoenzyme
Method: single particle / : Watson ER, Lander GC
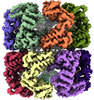
EMDB-45080: 
E.Faecium GroEL
Method: single particle / : Watson ER, Lander GC

EMDB-45098: 
E.coli GroEL + PBZ1587 inhibitor C1 reconstruction
Method: single particle / : Watson ER, Lander GC

EMDB-42116: 
HCN1 complex with propofol
Method: single particle / : Kim ED, Nimigean CM

EMDB-42117: 
HCN1 nanodisc
Method: single particle / : Kim ED, Nimigean CM
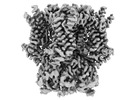
EMDB-44425: 
HCN1 M305L with propofol
Method: single particle / : Kim ED, Nimigean CM

EMDB-44426: 
HCN1 M305L holo
Method: single particle / : Kim ED, Nimigean CM

PDB-8uc7: 
HCN1 complex with propofol
Method: single particle / : Kim ED, Nimigean CM

PDB-8uc8: 
HCN1 nanodisc
Method: single particle / : Kim ED, Nimigean CM

PDB-9bc6: 
HCN1 M305L with propofol
Method: single particle / : Kim ED, Nimigean CM

PDB-9bc7: 
HCN1 M305L holo
Method: single particle / : Kim ED, Nimigean CM

EMDB-43746: 
Plasmodium falciparum 20S proteasome bound to an inhibitor
Method: single particle / : Han Y, Deng X, Ray S, Chen Z, Phillips M

PDB-8w2f: 
Plasmodium falciparum 20S proteasome bound to an inhibitor
Method: single particle / : Han Y, Deng X, Ray S, Chen Z, Phillips M
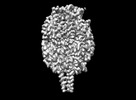
EMDB-43516: 
Cryo-EM structure of HMPV (MPV-2cREKR)
Method: single particle / : Yu X, Langedijk JPM

EMDB-43517: 
Cryo-EM structure of HMPV (MPV-2cREKR)
Method: single particle / : Yu X, Langedijk JPM

PDB-8vt2: 
cryo-EM structure of HMPV (MPV-2c)
Method: single particle / : Yu X, Langedijk JPM

PDB-8vt3: 
cryo-EM structure of HMPV (MPV-2cREKR)
Method: single particle / : Yu X, Langedijk JPM

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38874: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38875: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search






 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
